Portable genomics sheds light on hidden dynamics of last-resort antibiotic resistance
Antimicrobial resistance is one of the most pressing public health challenges of our time. This study uses portable nanopore sequencing to uncover hidden transmission routes of last-resort antibiotic resistance in a hospital setting, moving genomics from retrospective research tool to everyday clinical instrument.
Antimicrobial resistance is a long-standing and escalating global health challenge. Bacterial AMR was directly responsible for an estimated 1.27 million deaths worldwide in 2019, and the trajectory is worsening: a 2024 Lancetstudy projects that AMR will cause 39 million deaths between 2025 and 2050. Yet routine diagnostic tests reveal only that resistance exists, not how it spreads; and crucially, they lack the genetic detail to identify the exact mechanism of resistance or the specific bacterial strain involved, making it difficult to determine with confidence whether transmission has actually occurred. This study helps close that gap by using nanopore sequencing technology to follow resistance-carrying genetic elements in bacteria from both patients and hospital drains, uncovering hidden reservoirs and transmission routes that would otherwise stay undetected. With a workflow that can run on a standard computer directly in the lab, the team of researchers, including members of the Helmholtz AI Urban Group at Helmholtz Munich, shows how genomics can become a practical, day-to-day tool for infection control rather than just a retrospective research method. In the long term, these insights could help hospitals act earlier and more precisely, focusing on specific rooms, drains, and high-risk bacterial strains to slow the spread of multidrug-resistant pathogens and better protect vulnerable patients.
At a glance
In collaboration with clinical partners at the Institute of Medical Microbiology, Immunology and Hygiene at the Technical University of Munich, a research team including Ela Sauerborn - a physician and PhD student at Helmholtz AI - Daniel Gygax, and Lara Urban from the Helmholtz AI Urban Group at Helmholtz Munich has demonstrated how portable long-read nanopore sequencing can resolve the complex spread of plasmid-encoded carbapenem resistance in a hospital ward. The research article, titled "Resolving plasmid-encoded carbapenem resistance dynamics and reservoirs in a hospital setting through nanopore sequencing," was published in Microbial Genomics, Volume 12, Issue 2.
Why it matters
Carbapenem-resistant Enterobacterales (CRE) are a major clinical threat because they are resistant to last-resort antibiotics, and their resistance genes often sit on mobile plasmids that can jump between species. Routine diagnostics can flag resistant isolates, but they rarely reveal how resistance emerges, persists, and spreads over time and between patients and the ward environment. By combining genomics and AI-ready analysis pipelines, the Urban group shows how hospitals can move towards real-time, genomics-informed infection prevention and control.
Research
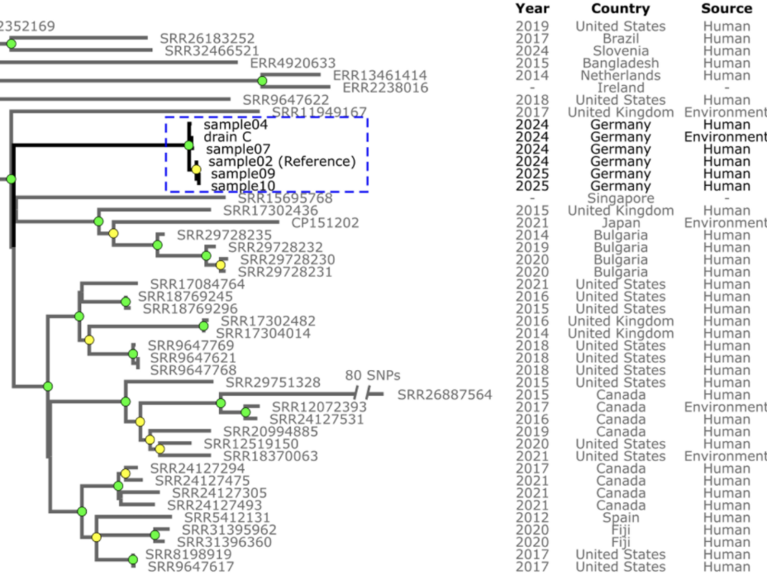
Over 13 months, the team retrospectively analysed 13 carbapenem-resistant Citrobacter isolates from ten patients and three sink/shower drains at the TUM University Hospital. Using Oxford Nanopore Technology's long-read sequencing and a laptop-based analysis workflow, they generated near-complete de novo assemblies of both bacterial chromosomes and carbapenemase-encoding plasmids, enabling high-resolution strain typing, plasmid profiling, and resistance gene detection directly from routine diagnostic isolates. Their pipeline integrates tools such as Dorado, Flye, Medaka, Pathogenwatch, AMRFinderPlus, MOB-suite, and Mash to provide an end-to-end, portable solution suitable for clinical and environmental samples.
One Health and AI perspective
The work exemplifies the Urban team's One Health vision of combining molecular biology and AI to understand and mitigate pathogen spread in complex ecosystems, from hospitals to the wider environment. This will allow AI-supported, real-time genomic surveillance that can help hospitals rapidly identify high-risk clones and plasmids and target infection prevention and control measures more precisely.